resistor
Module: devtests.bidomain.resistor.run
Section author: Gernot Plank <gernot.plank@medunigraz.at>
Validation of physical units used in extracellular voltage and current stimulation code
Problem Setup
This test generates a simle slab geometry of length 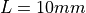 with a square cross section
of edge length
with a square cross section
of edge length 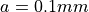 . A conductivity of
. A conductivity of 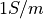 is assigned.
The total resistance between the two terminals of size
is assigned.
The total resistance between the two terminals of size 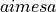 is then
is then 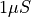 .
To use the extracellular space of the bidoman model as an approximation of a resistor,
a PASSIVE ionic model is used. As we aim to prevent current entering an intracellular path
we minimize current flow into the intracellular space by increasing the membrane resistance
and decreasing the conductivity of the intracellular current path.
.
To use the extracellular space of the bidoman model as an approximation of a resistor,
a PASSIVE ionic model is used. As we aim to prevent current entering an intracellular path
we minimize current flow into the intracellular space by increasing the membrane resistance
and decreasing the conductivity of the intracellular current path.
Two tests are performed. In both tests, the right hand side terminal will be grounded.
Into the left hand side terminal we either inject current of a total strength of 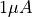 or we impose a voltage of
or we impose a voltage of 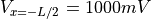 .
As we model a resistor in both cases the total voltage drop across the resisor has to be
.
As we model a resistor in both cases the total voltage drop across the resisor has to be 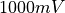 and the total current flowing over the resistor has to be
and the total current flowing over the resistor has to be 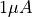 .
The setup is illustrated below:
.
The setup is illustrated below:

Usage
This example specifies only one argument, --stimulus,
which can be set to extra_V or extra_I.
./run.py --stimulus extra_I
Tests
extra_V
Checks:
- Compare against stored reference: max_error(phie.igb.gz)
- Compare against stored reference: max_error(vm.igb.gz)
Last run: 2024-02-29 01:04:04.045926, revision {‘base’: ‘cbf8efd0’}, dependency revisions {PT_C: 31642c1e,cvsys: 593686bc,eikonal: 5fbbfda3,elasticity: 4d92ddfc}
Runtime: 0:00:00.661865
ALL PASSED
PASS max_error(phie.igb.gz): 6.103515625e-05
PASS max_error(vm.igb.gz): 6.103515625e-05
extra_I
Checks:
- Compare against stored reference: max_error(phie.igb.gz)
- Compare against stored reference: max_error(vm.igb.gz)
Last run: 2024-02-29 01:04:04.753331, revision {‘base’: ‘cbf8efd0’}, dependency revisions {PT_C: 31642c1e,cvsys: 593686bc,eikonal: 5fbbfda3,elasticity: 4d92ddfc}
Runtime: 0:00:00.646867
ALL PASSED
PASS max_error(phie.igb.gz): 6.103515625e-05
PASS max_error(vm.igb.gz): 6.103515625e-05